批量计算基因和基因之间的相关性,也是一项很好的应用。
场景
画出某个基因和免疫细胞的相关性图
数据准备
数据准备
所需要的两项数据,一个是基因表达数据,另一个是免疫细胞浸润的矩阵,行名要一致,都是样本名称
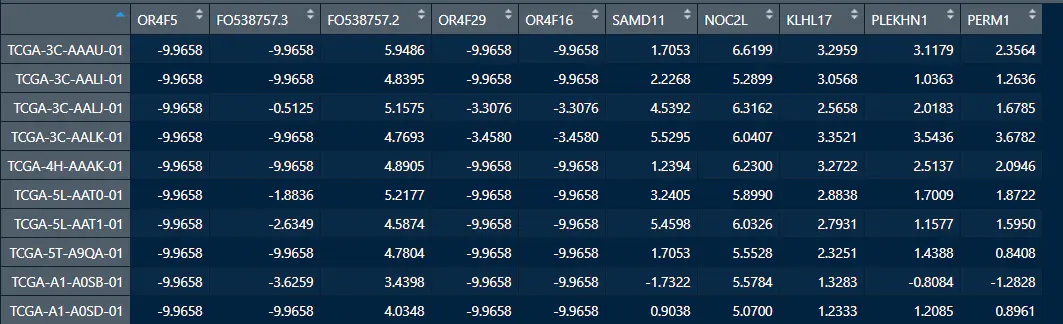
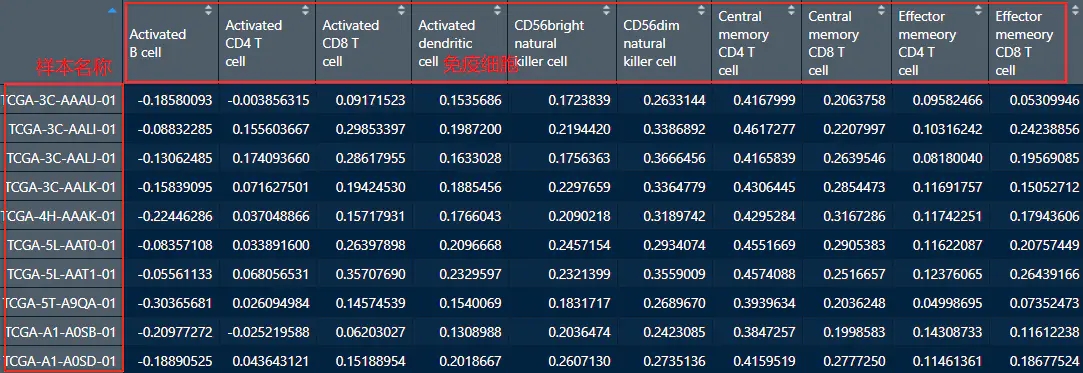
1. 准备画图函数
之前在批量处理——基因与免疫细胞的相关性已经计算过相关性,此次主要作图。
画图之前先对数据进行合并
plot_df <- cbind(immu_data, expr_data)写一个画图函数
ggcorplot <- function(a,b,method="spearman"){
corr_eqn <- function(x,y,digits=3) {
test <- cor.test(x,y,method=method,exact=FALSE)
paste(paste0("n = ",length(x)),
paste0("r = ",round(test$estimate,digits),"(",method,")"),
paste0("p value ",ifelse(test$p.value>0.001,paste0("= ",round(test$p.value,digits)),"< 0.001")),
sep = ", ")
}
plot_df <- plot_df[,c(a,b)]
names(plot_df) <- c("geneA","geneB")
require(ggplot2)
ggplot(plot_df,aes(geneA,geneB))+
geom_point(col="black")+
geom_smooth(method=lm, se=T,na.rm=T, fullrange=T,size=2,col="red")+
geom_rug(col="#006fbc")+
theme_minimal()+
xlab(paste0(a," relative expression"))+
ylab(paste0(b," relative score"))+
## 依靠函数来生成title
labs(title = corr_eqn(plot_df$geneA,plot_df$geneB))+
theme(plot.title = element_text(hjust = 0.5),
plot.margin = margin(1, 1, 1, 1, "cm"))
}这个画图函数是借鉴果子学生信课程
2.检验画图函数
yourgene = 'DDR1'
ggcorplot(yourgene, colnames(immu_data)[1], method = 'spearman')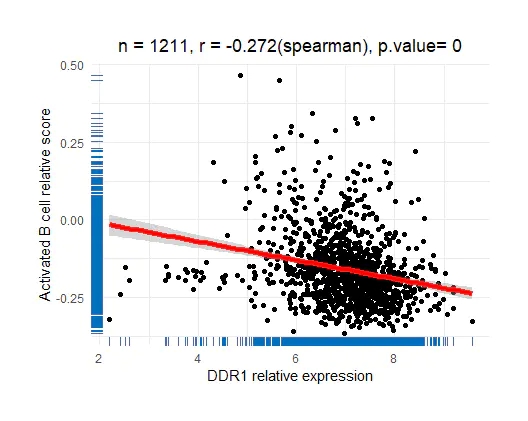
效果还可以
3. 批量处理
有了函数,直接套用批量处理——for循环画图的步骤就可以得到图
picDir <- './cor_picture/'
if (!dir.exists(picDir)) {
dir.create(picDir)
}
yourgene = 'DDR1'
for (i in colnames(immu_data)) {
print(i)
inputgene = yourgene
p = ggcorplot(yourgene, i, method = 'spearman')
ggsave(p, filename = paste0(picDir, inputgene, '_',i,'_correlation', '.pdf'), width = 7, height = 5)
}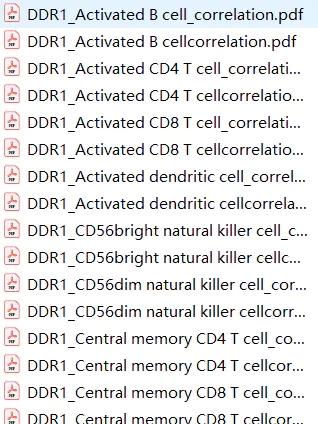
批量出图效果还是不错的。
4. 完整代码
plot_df <- cbind(immu_data, expr_data)
ggcorplot <- function(a,b,method="spearman"){
corr_eqn <- function(x,y,digits=3) {
test <- cor.test(x,y,method=method,exact=FALSE)
paste(paste0("n = ",length(x)),
paste0("r = ",round(test$estimate,digits),"(",method,")"),
paste0("p.value= ",round(test$p.value,digits)),
sep = ", ")
}
plot_df <- plot_df[,c(a,b)]
names(plot_df) <- c("geneA","geneB")
require(ggplot2)
ggplot(plot_df,aes(geneA,geneB))+
geom_point(col="black")+
geom_smooth(method=lm, se=T,na.rm=T, fullrange=T,size=2,col="red")+
geom_rug(col="#006fbc")+
theme_minimal()+
xlab(paste0(a," relative expression"))+
ylab(paste0(b," relative score"))+
## 依靠函数来生成title
labs(title = corr_eqn(plot_df$geneA,plot_df$geneB))+
theme(plot.title = element_text(hjust = 0.5),
plot.margin = margin(1, 1, 1, 1, "cm"))
}
ggcorplot(yourgene, colnames(immu_data)[1], method = 'spearman')
picDir <- './cor_picture/'
if (!dir.exists(picDir)) {
dir.create(picDir)
}
for (i in colnames(immu_data)) {
print(i)
inputgene = yourgene
p = ggcorplot(yourgene, i, method = 'spearman')
ggsave(p, filename = paste0(picDir, inputgene, '_',i,'_correlation', '.pdf'), width = 7, height = 5)
}
已收到回复,感谢你的帮助,祝你身体健康,工作顺利!
你好,请问为何图中P值会显示为0呢,是P值太小的原因吗。
修改一下自定义函数里面的这句话 paste0(“p.value= “,round(test$p.value,digits)),
我已修改 请参考 paste0(“p value “,ifelse(test$p.value>0.001,paste0(“= “,round(test$p.value,digits)),”< 0.001"))