require("survival")
fit<- survfit(Surv(time, status) ~ sex, data = lung)
# Basic survival curves
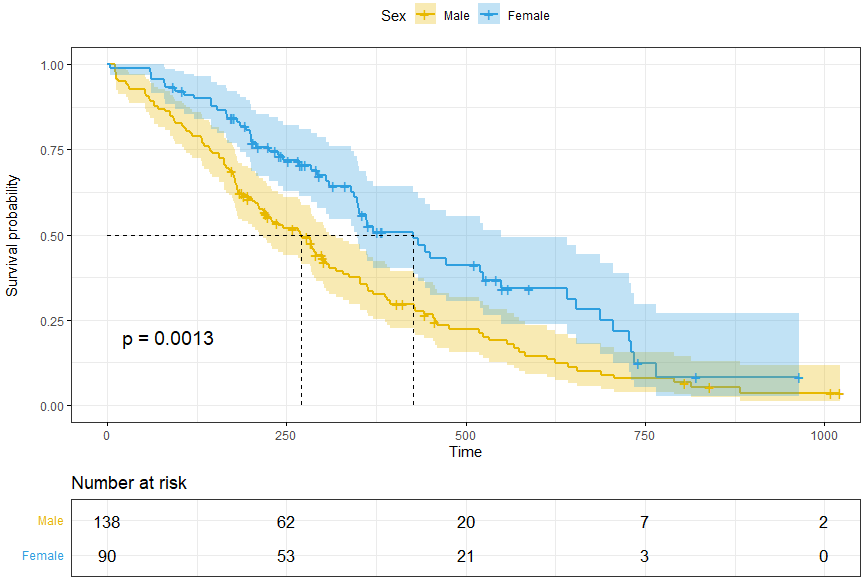
ggsurvplot(fit, data = lung)
# Customized survival curves
p <- ggsurvplot(fit, data = lung,
surv.median.line = "hv", # Add medians survival
# Change legends: title & labels
legend.title = "Sex",
legend.labs = c("Male", "Female"),
# Add p-value and tervals
pval = TRUE,
conf.int = TRUE,
# Add risk table
risk.table = TRUE,
tables.height = 0.2,
tables.theme = theme_cleantable(),
# Color palettes. Use custom color: c("#E7B800", "#2E9FDF"),
# or brewer color (e.g.: "Dark2"), or ggsci color (e.g.: "jco")
palette = c("#E7B800", "#2E9FDF"),
ggtheme = theme_bw() # Change ggplot2 theme
)
p <- ggsurvplot(fit, data = lung,
surv.median.line = "hv", # Add medians survival
# Change legends: title & labels
legend.title = "Sex",
legend.labs = c("Male", "Female"),
# Add p-value and tervals
pval = F, ##不默认显示P
conf.int = TRUE,
# Add risk table
risk.table = TRUE,
tables.height = 0.2,
tables.theme = theme_cleantable(),
# Color palettes. Use custom color: c("#E7B800", "#2E9FDF"),
# or brewer color (e.g.: "Dark2"), or ggsci color (e.g.: "jco")
palette = c("#E7B800", "#2E9FDF"),
ggtheme = theme_bw() # Change ggplot2 theme
)
##计算公式
data.survdiff <- survdiff(Surv(time, status) ~ sex,data = lung)
p.val = 1 - pchisq(data.survdiff$chisq, length(data.survdiff$n) - 1)
HR = (data.survdiff$obs[2]/data.survdiff$exp[2])/(data.survdiff$obs[1]/data.survdiff$exp[1])
up95 = exp(log(HR) + qnorm(0.975)*sqrt(1/data.survdiff$exp[2]+1/data.survdiff$exp[1]))
low95 = exp(log(HR) - qnorm(0.975)*sqrt(1/data.survdiff$exp[2]+1/data.survdiff$exp[1]))
ci <- paste0(sprintf("%.3f",HR)," [",sprintf("%.3f",low95),", ",sprintf("%.3f",up95),"]")
p$plot <- p$plot +
annotate("text", x = 0, y = 0.05,
label = paste0("P value = ",sprintf("%.3f",p.val),"\n HR (95% CI) = ",ci), ###添加P和HR 95%CI
size = 5, color = "black", hjust = 0)+
theme(text = element_text(size = 15))
p$plot


